
Ucsf Chimerax Structure Visualization For Researchers Educators And Chimera tutorials index structure analysis and comparison tutorial. this tutorial includes binding site analysis and comparison of related structures by superposition and morphing. internet connectivity is required to fetch the structures 3w7f and 2zco. background and setup; distances, h bonds, contacts; angles, rotamers, clashes; surfaces and. Ucsf chimera is a program for the interactive visualization and analysis of molecular structures and related data, including density maps, trajectories, and sequence alignments. it is available free of charge for noncommercial use. commercial users, please see chimera commercial licensing. we encourage chimera users to try chimerax for much.

Ucsf Chimera Structure Analysis Youtube Analyzing and comparing structures with ucsf chimera. structure analysis: hydrogen bonds and contacts amino acid sidechain conformations properties (b factor, hydrophobicity, etc.) comparison: superimposing structures morphing between structures an ensemble = multiple sets of coordinates for the same set of atoms. Learn how to use some of the tools of ucsf chimera to analyze and explore a protein structure. Structure analysis and comparison, html slides, updated january 2021. contains click to execute links. atp synthase cryoem tutorial given at stanford slac workshop, january 2020. contains click to execute links. alpha synuclein fibrils chimerax tutorial for rocky mountain laboratories workshop. september 2019. A suite of tools within ucsf chimera [1] has been developed for studying sequence structure relationships and comparing related structures. common tasks in sequence structure work include: (a) displaying information from a sequence alignment on one or more corresponding structures, or displaying information from the structures on the alignment.

Ucsf Chimerax Structure Visualization And Analysis Structure analysis and comparison, html slides, updated january 2021. contains click to execute links. atp synthase cryoem tutorial given at stanford slac workshop, january 2020. contains click to execute links. alpha synuclein fibrils chimerax tutorial for rocky mountain laboratories workshop. september 2019. A suite of tools within ucsf chimera [1] has been developed for studying sequence structure relationships and comparing related structures. common tasks in sequence structure work include: (a) displaying information from a sequence alignment on one or more corresponding structures, or displaying information from the structures on the alignment. Chimera tutorials index structure analysis and comparison tutorial. this tutorial includes binding site analysis and comparison of related structures by superposition and morphing. internet connectivity is required to fetch the structures 2zcp and 2zco. background and setup; distances, h bonds, contacts; angles, rotamers, clashes; surfaces and. Results: the molecular graphics program ucsf chimera includes a suite of tools for interactive analyses of sequences and structures. structures automatically associate with sequences in imported alignments, allowing many kinds of crosstalk. a novel method is provided to superimpose structures in the absence of a pre existing sequence alignment.
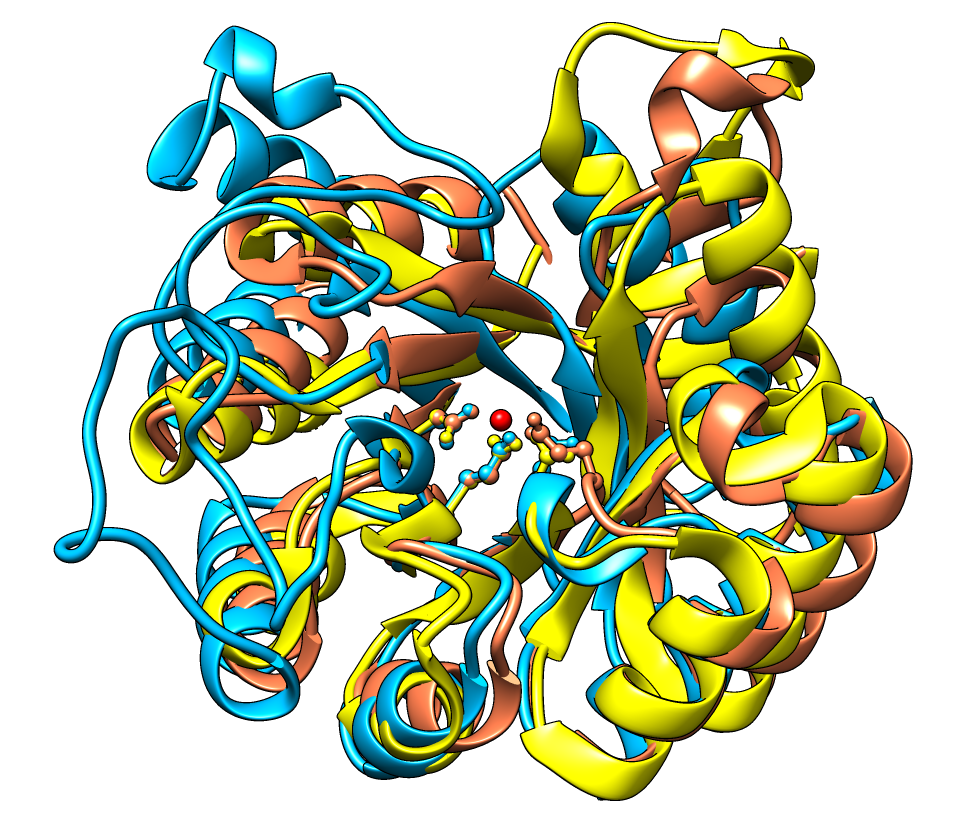
Ucsf Chimera Feature Highlights Chimera tutorials index structure analysis and comparison tutorial. this tutorial includes binding site analysis and comparison of related structures by superposition and morphing. internet connectivity is required to fetch the structures 2zcp and 2zco. background and setup; distances, h bonds, contacts; angles, rotamers, clashes; surfaces and. Results: the molecular graphics program ucsf chimera includes a suite of tools for interactive analyses of sequences and structures. structures automatically associate with sequences in imported alignments, allowing many kinds of crosstalk. a novel method is provided to superimpose structures in the absence of a pre existing sequence alignment.

Rs9616915 I245t Snp Structure Analysis By Ucsf Chimera H Bonding
