
Tutorial 1 вђ Getting Started With Ucsf Chimera Youtube This tutorial series will enable viewers to become proficient in using ucsf chimera ( cgl.ucsf.edu chimera ), an extensible molecular modeling sof. Ucsf chimera getting started. dna helix with bound netropsin. this tutorial provides an overview of basic features in chimera. you can interact with chimera using menus and or commands. the basic features of chimera are available either way, but not all command functions are available in menus or graphical interfaces, and not all menu or.
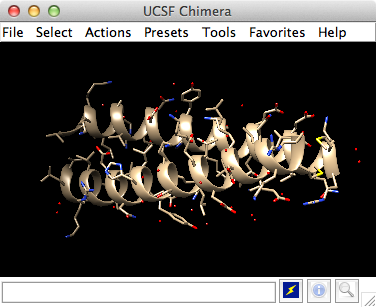
Getting Started With Ucsf Chimera Ucsf chimera tutorials. chimera tutorials. a set of tutorials is included in the chimera user's guide. the expanded "getting started" tutorial is more suitable for printing (more self contained rather than hyperlinked) than the above. video tutorials and tutorials from past chimera workshops are also available. It is necessary for running chimera on the mac. commands are entered into the command line and scaling and clipping operations can be performed with the side view. there are several ways to start each of these tools; one is to choose tools→keyboard→command line and tools→viewing parameters→side view from the menu. Commands, part 1 manipulation, selection, and chains. on windows mac, click the chimera icon; on unix, start chimera from the system prompt: unix: chimera. a basic chimera window should appear after a few seconds. commands are entered into the command line and scaling and clipping operations can be performed with the side view. Morphing attributes part 1 part 2 sequences structures alignments setup different proteins same protein model panel ensembles trajectories ensembles part 1 part 2 viewdock. more (web) help sheets. chimera quick ref (pdf) intro to pdb format.
Ucsf Chimera ж ќдѕње ґй ёгђђ1гђ й жіђиџў Yumuhan и з ѕ3dз жћ еџїи ењ е е е е и йў Commands, part 1 manipulation, selection, and chains. on windows mac, click the chimera icon; on unix, start chimera from the system prompt: unix: chimera. a basic chimera window should appear after a few seconds. commands are entered into the command line and scaling and clipping operations can be performed with the side view. Morphing attributes part 1 part 2 sequences structures alignments setup different proteins same protein model panel ensembles trajectories ensembles part 1 part 2 viewdock. more (web) help sheets. chimera quick ref (pdf) intro to pdb format. Chimera tutorials index getting started tutorial command version. many tasks in chimera can be accomplished in multiple ways. for example, colors and display styles can be changed with the actions menu or by entering commands. in general, commands are much more concise and powerful, but menus allow easy access to features without requiring. This command tells chimera to display the full amino acid structure, side chains and main chain, for residue 66 (: denotes residue, go to chimera getting started cmd for a cheatsheet). you can also use the dropdown menus for the same functions: select residue thr203 either by mousing over residues until you find thr203 or by adjusting the.
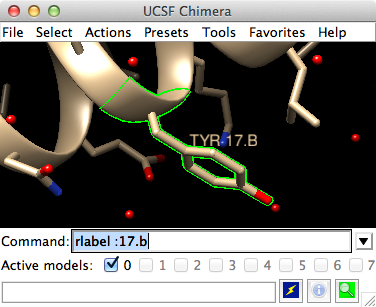
Getting Started With Ucsf Chimera Chimera tutorials index getting started tutorial command version. many tasks in chimera can be accomplished in multiple ways. for example, colors and display styles can be changed with the actions menu or by entering commands. in general, commands are much more concise and powerful, but menus allow easy access to features without requiring. This command tells chimera to display the full amino acid structure, side chains and main chain, for residue 66 (: denotes residue, go to chimera getting started cmd for a cheatsheet). you can also use the dropdown menus for the same functions: select residue thr203 either by mousing over residues until you find thr203 or by adjusting the.
